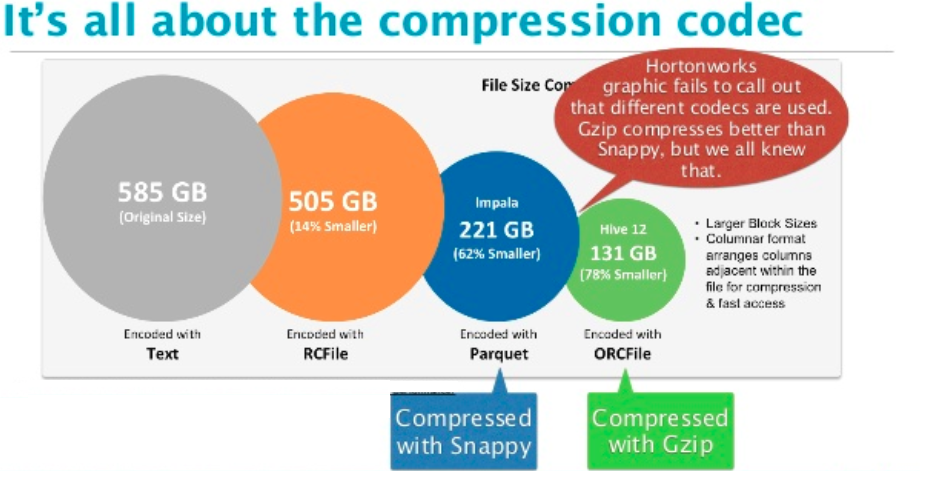
 
Compression definitionĭata compression is a technique that the main purpose is the reduction of the logical size of the file. The last section shows some compression tests. Only the second section lists and explains the compression codecs available in Parquet. However the first part recalls the basics and focuses on compression as a process independent of any framework. Next time when you need to store or quickly process large tables, consider Arrow/Parquet combination and save CSV file format for the final report.This post gives an insight on the compression provided by Apache Parquet. Working with large tabular files does not have to be a headache if done in a smart way. Parquet gives you metadata, quick way to check table dimensions without loading it, columns selection, it stores row indices, merging of partitions and much more… It later proved to be a great decision.īecause we switched to Parquet our analysis got almost order of magnitude faster when loading input files and then it go better. Parquet with Snappy compression – 2 threads, only first two columnsįor our calculation we were using only selected columns from our input files, but we wanted to preserve all of the original data. Parquet with Snappy compression – 2 loading threads Parquet with Snappy compression – 1 loading thread So the size difference is significant but not as significant as performance increase: Loading method 
Here are the file sizes for a small sample of our data set (~2 millions of rows): Format Please keep in mind that this is not a rigorous benchmark, just a crude comparison. To give you an idea of storage and performance benefits of using Parquet over text files let me show you some stats. # Aggregate the results into final report. Pool.apply_async(do_work, args=(filename,))ĭf = pq.read_table('out_dir', nthreads=mp.cpu_count()).to_pandas() Os.path.join('in_dir', 'iris-.parquet'.format(chunk_no)),Ĭhunk = pq.read_table(os.path.join('in_dir', filename), nthreads=2).to_pandas()įor filename in glob.glob('in_dir/*.parquet'): Reader = pd.read_table('', chunksize=1e7, names=column_names, compression='gzip')įor chunk_no, chunk in enumerate(reader): Split large, compressed file into Parquet chunks:Ĭolumn_names = To save time we decided to go with single server and Python’s multiprocessing module. Our dataset was a bit too large for a single machine but too small to bother with spinning a cluster. When paired with fast compression like Snappy from Google, Parquet provides a good balance of performance and the file size. To tame the input and the output files we used Apache Parquet, which is popular in Hadoop ecosystem and is the cool technology behind tools like Facebook’s Presto or Amazon Athena. It is backed by major players in every data processing ecosystem (Hadoop, Spark, Impala, Drill, Pandas, …) and supports a variety of popular programming languages: Java, Python, C/C++. The idea behind the project Arrow is to provide language-agnostic in-memory format for efficient handling of the columnar data. And it has a very simple API that plays nice with Pandas. Everybody knows there are no silver bullets but there is Apache Arrow. So we had to transform this into something more wieldy. Parallel processing of the large compressed files is next to impossible. That appeared to be a very smart choice until we had to parse this file a few times. 
What could we do with TSV that is larger than the size of local hard drives? The obvious approach was to just “gzipp” it. That needed to be parsed, filtered and aggregated into intermediate result for the next step of processing pipeline. For smaller datasets the solution would be trivial, but in this case about 5GB of input data grew to 2.5TB. Without going into much details, one of the steps of the analysis was a typical example of the large dataset vs a simple bioinformatics algorithms. Just recently we completed a very interesting project, which also provided us with a valuable lesson. When working with larger projects’ datasets sizes of hundreds of gigabytes or a few terabytes are not unusual. But once you start analysing this data you can get as much as 10 times the size of raw input data. Compressed output of a typical sequencing run is around few gigabytes. From software engineering point of view, when you skip fancy nomenclature and go beyond scientific paper abstracts, bioinformatics is mostly about handling tabular data. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
